Personalized Genome
Human genome is all the sequence information of human cell. It is used as a platform to describe primary structure of human genome. Completion of human genome project allows characterization of functional elements of human genome. Different versions of the same gene circulate within different group of people in different geographical groups. This phenomenon is called genetic polymorphism. Genetic polymorphism is reflected by sequence variation, such as single nucleotide polymorphisms (SNPs), copy number variations (CVs), insertions and deletions (InDels). Therefore a generalized human genome cannot be used to represent genome of different individual from different groups of people due to these genetic polymorphisms. As mentioned above, different versions of the same gene exist among human population. If “A” and “a” are two different versions of a given gene, we call them alleles. Human has diploid genome, meaning human cells contain two sets of genome, with one from mother and another one from father. If two alleles of a gene are the same (AA or aa), the genotype of this gene is called homozygote, otherwise heterozygote (Aa). If allele “a” is disease associated, heterozygosity of this gene or homozygosity for allele “a” poses a risk of the disease to certain individual.
Personalized genome is all the sequence information of one human individual. The individual-specific sequence differences (i.e. SNPs and InDels) can serve functions of interpreting ancestors’ ethical group and geographical origin and determining of kinship among individuals. More importantly, personalized genome serves as a biological feature that best represents one individual’s genome. The polymorphisms, and mutations, are major reasons for susceptibility to certain disease, and also the different-to-none response to medication. Judging on this, the value of personalized genome can provide a full detailed report of what disease one might be associated with, which environment risk factors can potentially contribute to certain health conditions, and finally facilitate medication selection for personalized medicine. With the technical advances of DNA/RNA sequencing technology, development of bioinformatics and data science, sequencing of personalized genome becomes a mature and affordable technique.
Personalized genome sequencing
Personalized genome sequencing strategies can be classified as genotyping microarray, exome sequencing and whole-genome sequencing.
Measuring genotypes of known disease related polymorphisms is capable of determining whether one is disease associated. Genotyping microarray is based on molecular hybridization between DNA/RNA sample of tester and pre-designed nucleotide probes. This method offers fast data turnover and low analysis cost. However the targets to be analyzed are limited by pre-designed probes based on known polymorphisms therefore this method cannot fully resolve variances within one’s genome.
Exome sequencing and whole-genome sequence provide the most comprehensive about one’s genome. Both sequencing strategies relies on Next Generation Sequencing (NGS) platforms. In addition to the information that can be acquired from genotyping analysis, exome sequencing includes sequencing information of all exons. Whole-genome sequence additionally includes all non-coding sequences, intergenic region sequences. Thus any sequence variation can be captured. 1000 genome project showed on average each individual carries 250-300 loss-of-function variants in known genes and 50 to 100 variants that have been implicated in inherited disorders. More comprehensive sequence information enhances the reorganization of these variants and therefore guarantees the accuracy of personalized medicine. DNA was usually sequenced at 30-50 fold of coverage to collect enough sequence information for statistically determining genetic variations. These variations can subsequently be compared to multiple databases to find associated disease.
Personalized medicine
Personalized medicine heavily relies on personalized genomes. For example, an advanced chemotherapy selection method developed in Mayo Clinic implements xenograft of patient derived tumor cells into immune-suppressed mouse. Tumor cell clones isolated from patients were amplified in multiple animals. Drug selection was subsequently performed on the tumor xenograft. Treatment effectiveness of a drug is determined on the effectiveness of the drug on repressing the tumor related gene expression profile shift in post-treatment cancer cells. Patient data suggested that genetic variations, gene expression and drug response are able to classify patients into different cohorts. Therefore using such cohorts, effective medication can be reversely estimated.
With the advance of genetic research technique and data science, availability of various types of data, sequencing cost is much lower compared to 4-5 years ago and is estimated less than 1,500 dollars. If you are interested to have your own genome sequenced and analyzed, feel free to contact with us for any question and we are happy to design most appropriate sequencing plan.
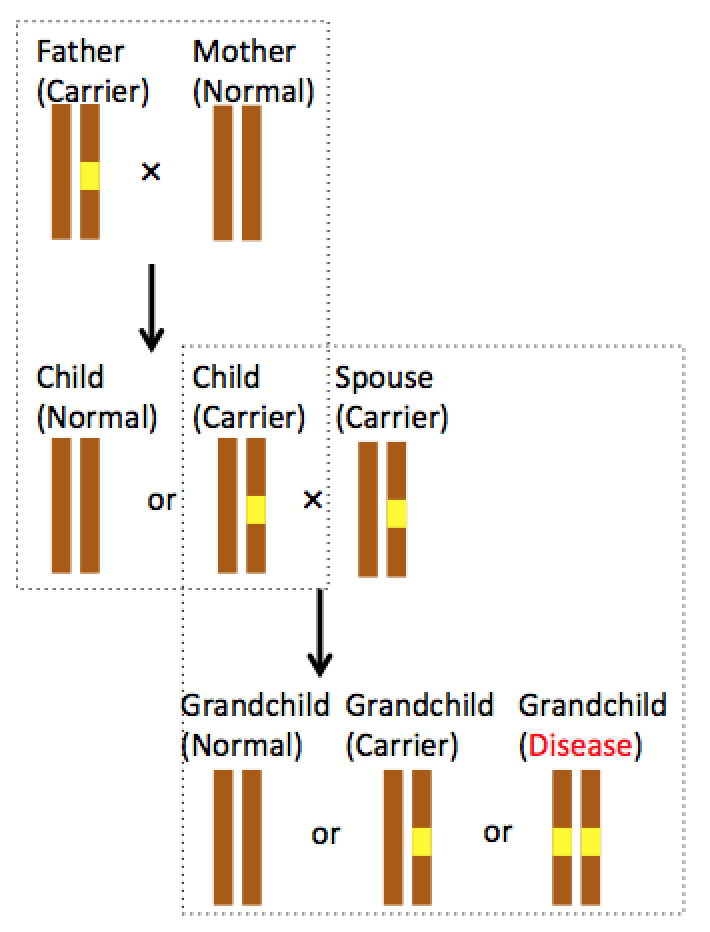
Figure 1 Disease related SNP
Figure 1 illustrates an example of how a recessive SNP cause disease. Assuming there are two different alleles in population for a given gene, one is not disease related (brown allele) and the other is disease related (yellow mixed allele). The disease related allele leads to disease in a recessive manner, meaning disease related allele produce its effect only when the gene is homozygous for disease related allele. Father carries one of each allele (heterozygosity) while mother carries only one allele (homozygosity). By Mendelian inheritance, each of their children has 50% of chance to carry the disease related allele. If the carrier child and another disease allele carrier have children, there are 25% of chance for their kids to be normal (homozygous for normal allele), 50% of chance to be carrier (heterozygous for the given gene) and 25% of chance to have the disease (homozygous for disease allele). If this disease has environmental risk factor(s), realizing one’s genome is prone to certain disease will be beneficial to avoid risk factor exposure and prevent disease from taking place, or lead to early preventative treatment.
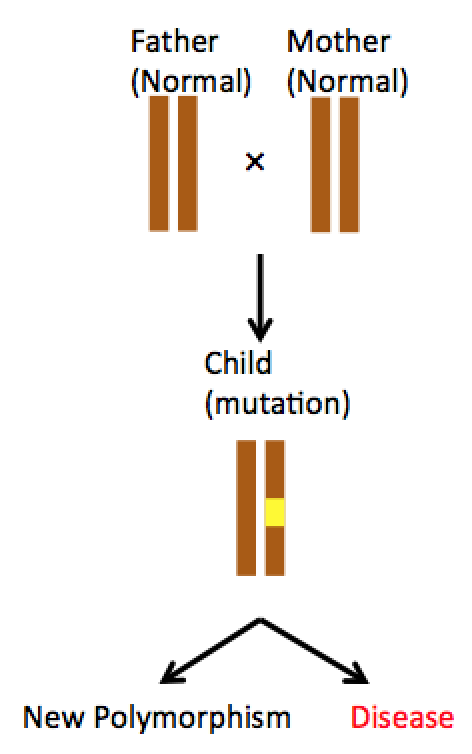
Figure 2 Mutation
Figure 2 illustrates an example of how a mutation cause disease. Mutations are not inherited from parents. Instead, mutation takes place in one’s own genome by different mechanisms.The bright side of a mutation is if it doesn’t have deteriorative effect, it can be passed onto next generation like a normal gene and circulate in population as a new polymorphism. On the other hand, not only detrimental mutations have damaged function, but also mutant’s gene product can inhibit normal allele’s product (i.e. tumor suppressor p53). In cancer research, Loss-of-heterozygosity (LOH) serves as a marker that indicates disease progression. Mutation originally took place in one of the two alleles, with the other normal allele can still produce normal gene products. When LOH is observed, both alleles are mutated, therefore the cell cannot produce normal gene products anymore.
个体基因组
个体基因组即个体的所有基因组核酸序列。随着人类基因组的全测序,以及测序技术,生物信息学和数据学的飞速发展,个体基因组的测序以及后续的数据分析已经是非常成熟的技术。个体基因组作为最符合受试者个体生物学特征的属性,可以作为依据来预测潜在疾病,并可以最大化地指导个体化医疗和个体化给药。
由于基因交流的限制(来源于生殖隔离,性选择隔离),不同的人类族群内部流传着不同版本的同一基因,这种现象称为基因多样性。基因多样性反映在基因序列的差异上,其表现为单核苷酸多样性(SNP),拷贝数差异(CNV)以及序列的插入或缺失(InDel)。人类基因组作为一个群体概念,它的序列信息仅作为一个参考来描述人类染色体的一级结构。因此人类基因组并不能代表不同族群以及族群中每一个个体的基因组。现有的医疗手段和药物并不能100%的治愈所有患者,其中一个很重要的原因正是个体基因组之间存在差异造成的。因此,个体基因组对于个人的健康监控以及疾病诊疗就有着重大意义。它不仅能通过种群学的数据推断受试者的种族,地域,判断多个个体之间是否存在亲缘关系,更重要的是能够为健康的受试者提供一份完备的基因组报告来显示受试者是否具有患特定疾病的可能,哪些环境因素会对受试者的健康产生影响,为患病的受试者提供潜在的或已知的对特定基因差异有效的治疗方法和药物。同一基因在不同人群中有不同的序列片段,假设基因A和基因a为同一基因在不同人群中的版本,A和a则称为等位基因。人类作为二倍体生物拥有两套基因组,一套来自父亲,一套来自母亲。如果两套基因组中的该基因都为A或a(AA或aa),我们称这种基因型为纯合子。反之如果一条基因是A,而另一条是a,这种基因型(Aa)则称为杂合子。如果等位基因a与遗传病相关,那么基因型为AA的个体无患病可能,Aa个体为致病基因携带者有患病可能,aa个体则患病。因此通过测定已知的疾病相关基因的SNP/InDel位点,分析基因型就能得知特定个体是否有患某种疾病的可能。
个体基因组的检测方法和意义
个体基因组有多种检测方式,取决于受试者的目的,可以分为基因型微矩阵分析,外显子测序,和全基因组测序。
基因型微矩阵分析是通过受试者生物样本和核酸探针之间的核酸分子杂交实现的。该方法检测速度快,数据分析简单,花费低。但是由于核酸探针局限于已设计的多样性位点,这种方法的结果并不全面。
外显子测序和全基因组测序提供了最完善的个体基因组信息。这两种发发都依赖于高通量测序的方法。这些数据信息不仅含括了基因型分析的结果,还在此基础上提供了所有编码以及非编码序列,SNP/Indel,拷贝数变化的信息。1000 human genome project(千人基因组计划)显示平均每个人携带了250到300个导致基因产物功能缺失的基因组序列差异,同时还携带了50到100个通过遗传获得的序列差异。更多的数据保证了对这些基因组序列差异以及他们相关的疾病辨识度和药物筛选的准确性。在进行全基因组测序时,我们一般会对基因组进行30-50X深度的测序,以保证在统计学上有足够多的序列信息来发现不同的等位基因和突变。而后我们会将这些数据与人类基因组参考数据比对并把序列差异与多个数据库比对来发现潜在的疾病信息。
个体化医疗依赖于这些个体化基因组序列信息。一种先进的个体化医疗方案就是利用病人肿瘤来源的癌细胞接种到免疫系统缺陷的小鼠体内得到多个同源肿瘤克隆,并在这些克隆中筛选药物来治疗病人的肿瘤。对于一种药物的有效性评价是通过药物治疗后肿瘤的基因表达是否更接近于正常组织来判定的。大量的病人数据显示基因表达相似的病人有着相似的基因序列特性,也对同一药物产生积极的治疗效果。利用几千份这样的公共数据和临床数据,受试者的个体基因组可以与他们比对并推算出有效的治疗药物。多种样品可以用来进行测序分析,比如唾液,血液,羊水,活组织检验样本,病理样本样品等。样本的易获取性使个体化基因组的测定可以与日常的健康检查,孕期检查,以及早期的疾病诊断相结合。
图1 疾病相关的单核苷酸多样性差异
图1展示了一个单核苷酸多样性是如何造成某种隐性病(注意,并不是多样性一定与疾病有关,很多情况下多样性只表现在表型的区别)。假设人群中某基因有两条等位基因,其中一条不引起疾病(棕色),另一条与疾病相关(棕色和黄色相间,黄色代表多样性)。这条疾病相关的等位基因引起隐性遗传病,所以只有在个体携带两条疾病相关的等位基因时才患病。家族中的父本基因组携带两种等位基因(杂合子)而母亲只携带正常的等位基因(纯合子)。根据孟德尔遗传规律,亲本双方的每个孩子都有50%的可能携带致病基因。如果携带致病基因的子女与另一个该致病基因携带者,那么该子女的下一代有25%的机会不懈怠致病基因,50%的机会携带致病基因但不患病,还有25%的机会患遗传病(致病基因纯合子)。如果该疾病有一个或多个环境危险因子,那么了解基因序列以及基因型对于遗传疾病的早期预防和治疗都是有利的。
图2 突变
图2显示了突变的发生。突变不能从亲本遗传获得,而是在个体中由有多重原因造成的。突变往往只发生在两条等位基因上的一条,在这种情况下机体有一条突变体基因和一条正常基因,但是突变不仅造成基因产物功能上的受损或缺失,突变基因的产物还会限制正常等位基因产物的功能(比如抑癌蛋白p53的突变产物和正常基因产物形成多聚体从而使正常基因产物失活)。在癌症研究中,突变基因的杂合性缺失(LOH)往往说明相对于突变基因的正常基因静息,使得机体不再表达任何正常的基因产物。LOH因此通常和疾病的恶化挂钩。对于突变的另一方面,如果突变不存在坡滑行的效果,那么它可以像一个正常基因一样被传播到子代基因组并作为一个新的基因多样性流传在人群中。
从基因研究的角度来讲我们生活在一个黄金年代。遗传学,基因组学和数据学的发展以及相关技术的革新,创造了大量的数据以及相比于4,5年前低廉的基因组测序价格。如果您对个体基因组感兴趣,欢迎联系我们,我们会针对您的需求设计最佳测序方案。
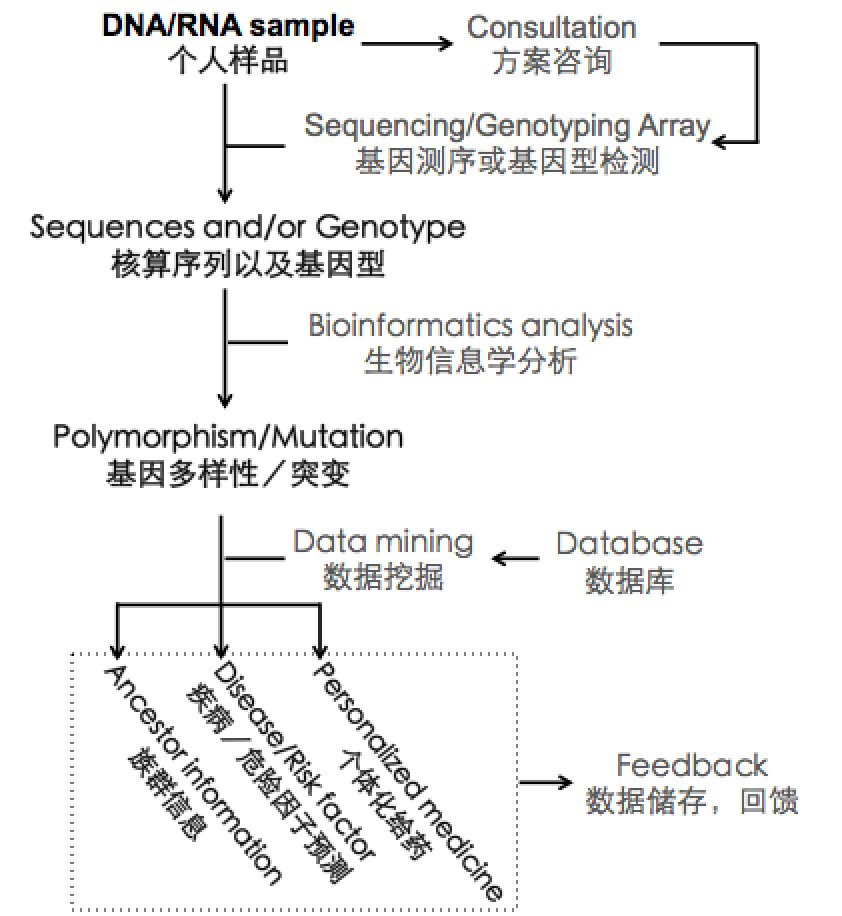
How to have personalized genome sequenced
If you are interested in sequencing your own genome, we recommend you initiate the whole process by contacting us. Based on your need, we can help you to design the best project protocol, arrange your sample collection and sequencing service. For example, if you are interested in where did you ancestor come from, we will recommend performing genotyping array plus population genetics analysis to give you the desired answer with low budget. In contrast, if you have the desire of fully sequencing your personal genome, a more complicated sequencing would be more appropriate. We will arrange your sample to be sequenced at our collaborating sequencing service provider that is close to you. After sequencing is accomplished, the sequencing data will be piped to our server for downstream analysis. Using state-of-art bioinformatics tools, we will determine gene polymorphisms and gene mutations. Subsequently, we will compare your genome to known genomic information to determine whether your genetic variation is associated with certain health condition. In the final report that will be send back to you, we will report any polymorphism and mutation we found in your genome, as well as any disease and risk factors that are potentially associated with them. We will also report any drug shown to repress the function of certain genetic variations. After the whole process, we will contact you again to go through details of your analysis result, and whether you want to share your genetic information with your health provider when necessary. If you are interested in a genome project about yourself, please contact with us.
如何检测个体化基因组
如果您希望检测自己的基因组序列,我们建议您先与我们联系。基于您的预期,我们可以帮助您设计最合适的项目流程,安排您的样品采集和测序服务。比如,如果您对您家族祖先的地理属性感兴趣,我们会推荐您做基因型分析的micro-array,以及种群遗传学的分析,而不推荐深度测序,因为前者足以满足您的需求而且价格相对便宜很多。但是如果您希望了解自己的基因组的每一个碱基序列以及所有的基因多样性,那么我们会推荐您做深度测序。在这之后,我们会安排您的基因组样品在就近的合作测序中心完成。测序结果会传送到我们的服务器上进行下游分析。我们会用最新的生物信息学工具来检测您基因组中的基因多样性和基因突变。接下来我们会将您基因组中发现的任何差异与已知的差异做比较来确定您基因组中的差异是否与特定的健康状态有关。在最终交给您的报告中,我们会列举所有检测到的基因多样性基因突变,以及这些基因组差异相关的疾病以及危险因子,同时我们也会报告有对抗致病突变活性的药物。在整个测序和分析过程完成之后,我们会再次与您联系。一方面我们会跟您解释所有的分析结果;另一方面,我们需要与您讨论事后需要将您的遗传信息分享给您的医生以方便更好的健康服务和监控。如果您对个体化基因组测序感兴趣,欢迎您与我们联系。